Day 26 Planning designed experiments
26.1 Announcements
- Project deadlines:
- December 12th: give your presentation by this date (it can be earlier than this also). You will need to schedule a 30min meeting with me.
- December 19th: submit your final project and tutorial on canvas, including both your classmate’s and my feedback.
26.2 Designed experiments
- On the importace of designed experiments
- Defining the data generating process
- Statistical power
- Precision of parameters
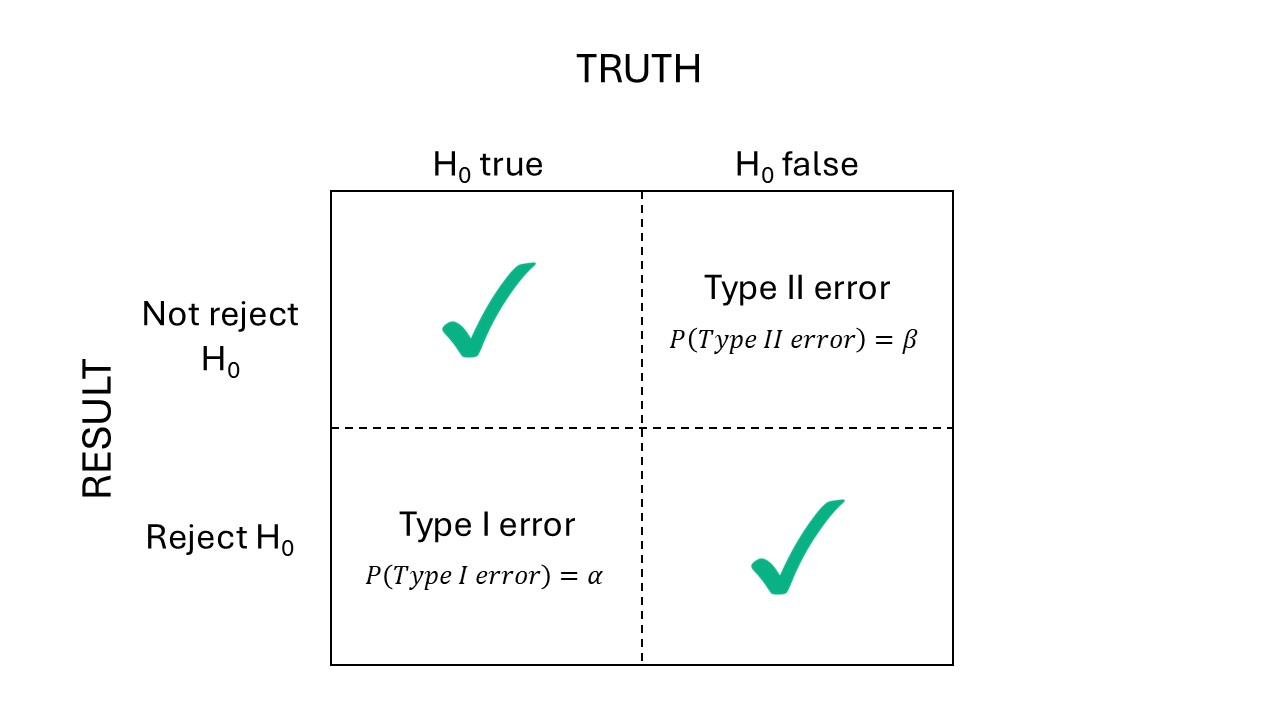
Figure 26.1: Types of errors.
26.2.0.1 Analytical solutions
We know Power is
\[Power = P(F>F_{c} \vert F \sim F(df_1, df_2, \phi)),\]
where:
- \(F\) is the F test statistics,
- \(F_c\) is the critical test F,
- \(df_1\) are the degrees of freedom of the numerator,
- \(df_2\) are the degrees of freedom of the denominator,
- \(\phi\) is the noncentrality parameter.
Note that \(F\) depends on \(\sigma^2_\varepsilon\), and \(F_c\) depends on \(\alpha\).
Suppose we are trying to compare designs to find differences main effects and interactions of a \(3 \times 2\) factorial treatment structure. Suppose we have:
- Treatment A: 3 levels,
- Treatment B: 2 levels.
Typically, we would find means around \(10\) units, and \(\sigma^2 \approx 1\).
This is how a dataframe coudl look like:
n_reps <- 3
data <-
purrr::map_dfr(
seq_len(n_reps),
~expand.grid(tA = factor(1:3),
tB = factor(1:2)) |>
mutate(mu = c(10,12,12,10,10,12))) |>
mutate(rep = rep(1:n_reps, each = 6))
head(data)## tA tB mu rep
## 1 1 1 10 1
## 2 2 1 12 1
## 3 3 1 12 1
## 4 1 2 10 1
## 5 2 2 10 1
## 6 3 2 12 1How do different designs compare?
# create an RCBD model
m_rcbd <- lme(mu ~ tA * tB,
random = list(rep = pdSymm(matrix(sqrt(sigma2_block)), ~ 1, nam = "(Intercept)")),
data = data,
# this won't converge but ask to get an object anyways
control = lmeControl(
sigma = sqrt(sigma2_e),
returnObject = TRUE))
# peek at the variance-covariance matrix
VarCorr(m_rcbd)## rep = pdSymm(1)
## Variance StdDev
## (Intercept) 0.00863157 0.09290624
## Residual 1.00000000 1.00000000# fix variance components
m_rcbd$modelStruct$reStruct$rep <- pdSymm(matrix(sqrt(sigma2_block)), ~ 1, nam = "(Intercept)")
# peek at the variance-covariance matrix again
VarCorr(m_rcbd)## rep = pdSymm(1)
## Variance StdDev
## (Intercept) 0.8944272 0.9457416
## Residual 1.0000000 1.0000000results <-
emmeans::joint_tests(m_rcbd) |>
as.data.frame() |>
transmute(source = `model term`,
numDF = df1,
denDF = df2,
F.ratio)
alpha <- 0.05
results |>
mutate(ncp = F.ratio* numDF,
Fcrit = qf(1-alpha, df1 = numDF, df2 = denDF, ncp = 0),
Power = 1 - pf(Fcrit, df1 = numDF, df2 = denDF, ncp = ncp))## source numDF denDF F.ratio ncp Fcrit Power
## 1 tA 2 10 6 12 4.102821 0.7592131
## 3 tB 1 10 2 2 4.964603 0.2490518
## 2 tA:tB 2 10 2 4 4.102821 0.3183418# create a split-plot model where tA was the whole plot
m_sp <- lme(mu ~ tA * tB,
random = list(rep = pdSymm(matrix(sqrt(sigma2_block)), ~ 1,
nam = "(Intercept)"),
tA = pdSymm(matrix(sqrt(sigma2_wp)), ~ 1)),
data = data,
# this won't converge but ask to get an object anyways
control = lmeControl(sigma = sqrt(sigma2_e),
returnObject = TRUE,
maxIter = 0))
# peek at the variance-covariance matrix
VarCorr(m_sp)## Variance StdDev
## rep = pdSymm(1)
## (Intercept) 0.01216669 0.1103027
## tA = pdSymm(1)
## (Intercept) 0.03296807 0.1815711
## Residual 1.00000000 1.0000000# fix variance components
m_sp$modelStruct$reStruct$rep <- pdSymm(matrix(sqrt(sigma2_block)), ~ 1, nam = "(Intercept)")
m_sp$modelStruct$reStruct$tA <- pdSymm(matrix(sqrt(sigma2_wp)), ~ 1, nam = "(Intercept)")
# peek at the variance-covariance matrix again
VarCorr(m_sp)## Variance StdDev
## rep = pdSymm(1)
## (Intercept) 0.8944272 0.9457416
## tA = pdSymm(1)
## (Intercept) 0.4472136 0.6687403
## Residual 1.0000000 1.0000000results <-
emmeans::joint_tests(m_sp) |>
as.data.frame() |>
transmute(source = `model term`,
numDF = df1,
denDF = df2,
F.ratio)
alpha <- 0.05
results |>
mutate(ncp = F.ratio* numDF,
Fcrit = qf(1-alpha, df1 = numDF, df2 = denDF, ncp = 0),
Power = 1 - pf(Fcrit, df1 = numDF, df2 = denDF, ncp = ncp))## source numDF denDF F.ratio ncp Fcrit Power
## 1 tA 2 4 5.629 11.258 6.944272 0.5146767
## 3 tB 1 6 2.000 2.000 5.987378 0.2231880
## 2 tA:tB 2 6 2.000 4.000 5.143253 0.268217926.2.1 Precision analysis
- Power analysis versus precision analyses (the focus on p-values)
- ASA’s statement on p-values
- Scientists rise up against statistical significance
- Discussion
- Counterargument: In defense of p-values
- Gelman “Let us have the serenity to embrace the variation that we cannot reduce, the courage to reduce the variation we cannot embrace, and the wisdom to distinguish one from the other.” [see talk]
## NOTE: Results may be misleading due to involvement in interactions## [1] 2.572833## NOTE: Results may be misleading due to involvement in interactions## [1] 3.3099726.3 Bayesian analysis of designed experiments
Bayes theorem:
\[[\boldsymbol{\theta} \vert \mathbf{y}] = \frac{[\mathbf{y} \vert \boldsymbol{\theta}] [\boldsymbol{\theta}] }{ [\mathbf{y}]}, \]
where:
- \([\boldsymbol{\theta} \vert \mathbf{y}]\) is the posterior distribution,
- \([\mathbf{y} \vert \boldsymbol{\theta}]\) is the likelihood,
- \([\boldsymbol{\theta}]\) is the prior,
- \([\mathbf{y}]\) is the data.
Note that whatever is defined in \([\cdot]\), it’s information.
26.4 Applied example
Suppose we designed an experiment with the rationale outlined above. What if the model doesn’t converge?
- About dropping effects from the model
- Bayesian solutions
dat <- readxl::read_xlsx("../../data_confid/Behavior_Counts.xlsx")
# viz
dat |>
ggplot(aes(timepoint, behavior_count))+
geom_line(aes(group = calfID,
color = breed))+
geom_point(aes(group = breed,
fill = breed),
shape =21, size= 2.5)+
facet_wrap(~trt)+
scico::scale_fill_scico_d()+
scico::scale_color_scico_d()+
theme_pubclean()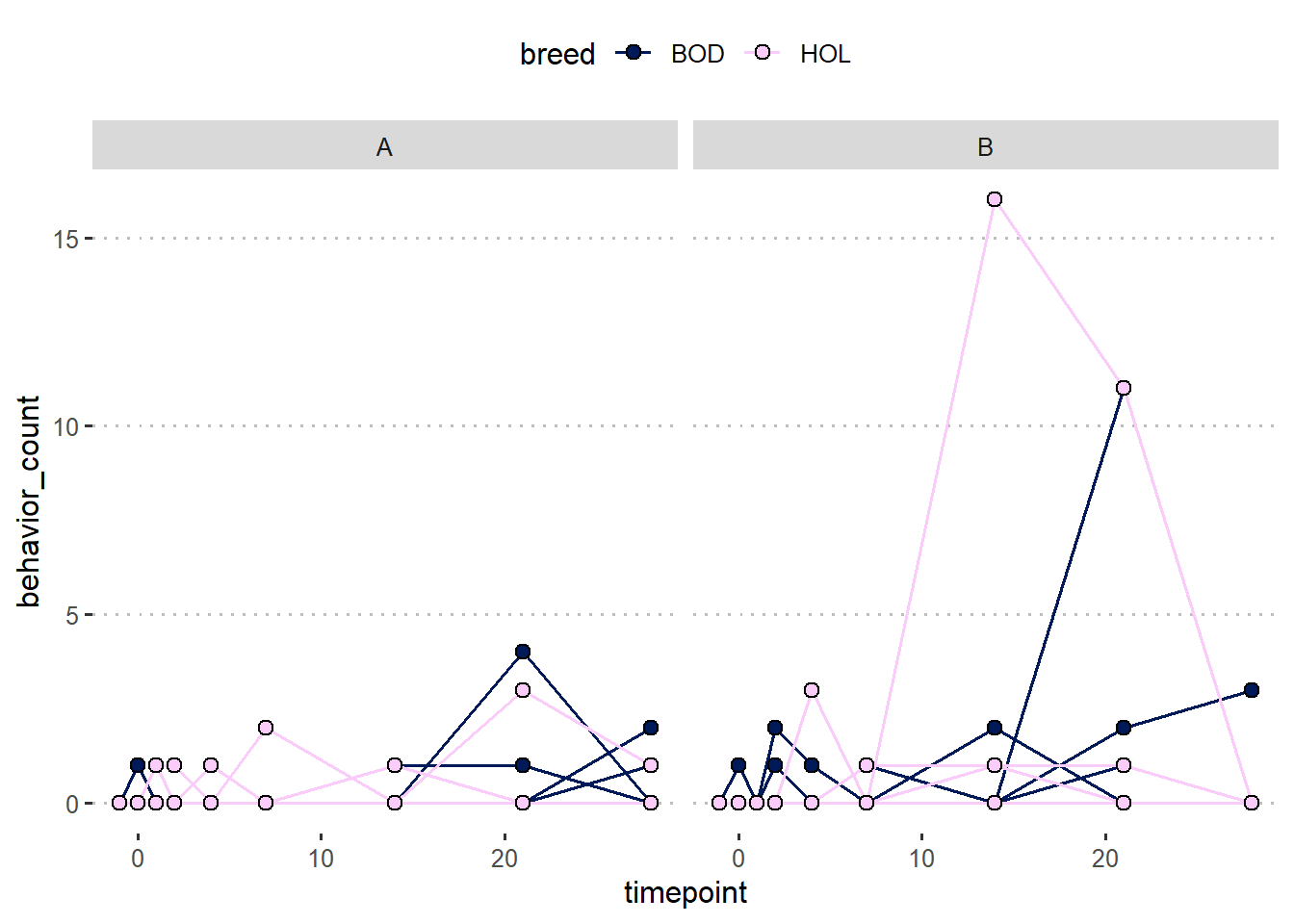
m2 <- glmmTMB(behavior_count ~ timepoint * trt * breed +
toep(timepoint + 0 | calfID),
ziformula = ~1,
family = poisson(link = "log"),
data = dat |> mutate(timepoint_f=as.factor(timepoint)))What to do?