Day 13 Modeling designed experiments in the presence of spatial variability
October 8th
13.2 Spatial variability
- Most commonly found in field experiments.
- Whiteboard: going from the normal iid model to a spatial dependence model.
13.2.1 Spatial patterns
Consider the model \[\mathbf{y} \sim N(\boldsymbol{\mu}, \boldsymbol{\Sigma}),\] where \(\boldsymbol{\Sigma}\) is the variance-covariance model. In the presence of spatial patterns, there are parametric and non-parametric tools to account for spatial dependence.
Sometimes, we can clearly define the shape of the spatial dependence pattern. For example:
Matérn covariance function
\[C(d; \sigma^2, \phi, \kappa) = \sigma^2 \frac{(h/\phi)^\kappa K_\kappa (h/\phi)}{2^{\kappa-1} \Gamma(\kappa)}\]
Gaussian covariance function
\[C(d; \sigma^2, \phi) = \sigma^2 \exp \left\{ - \frac{d^2}{2\phi^2} \right\} \]
Exponential covariance function
\[C(d; \sigma^2, \phi, \kappa) = \sigma^2 \exp\{-(d/\phi)^\kappa\}\]
13.2.2 Applied example
Yield data from an advanced Nebraska Intrastate Nursery (NIN) breeding trial conducted at Alliance, Nebraska, in 1988/89.
library(tidyverse)
library(agridat)
library(ggpubr)
data <- drop_na(stroup.nin)
data %>%
ggplot(aes(col, row))+
geom_tile(aes(fill = rep), color = "black")+
theme_pubr()+
labs(fill = "Block")+
theme(axis.line = element_blank())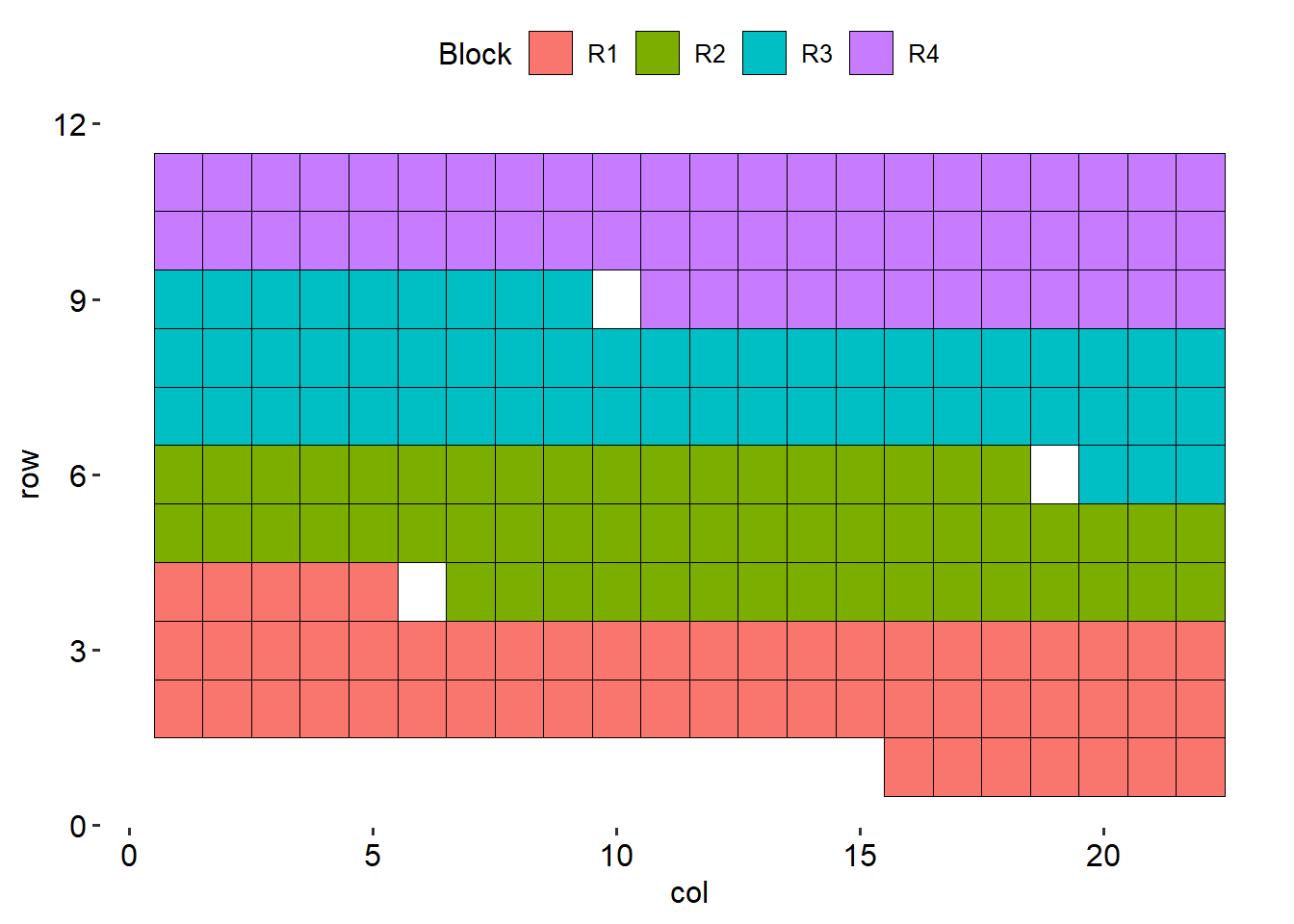
The statistical model:
\[y_{ij}\sim N(\mu_{ij}, \sigma^2), \\ \mu_{ij} = \eta_{ij} = \eta_0 + G_i + b_j, \\ b_j \sim N(0, \sigma^2_b).\]
Assumptions:
- iid, normal residuals
data %>%
ggplot(aes(col, row))+
geom_tile(aes(fill = res), color = "black")+
theme_pubr()+
scico::scale_fill_scico()+
theme(axis.line = element_blank())+
labs(fill = expression(Residual~(bu~ac^{-1})))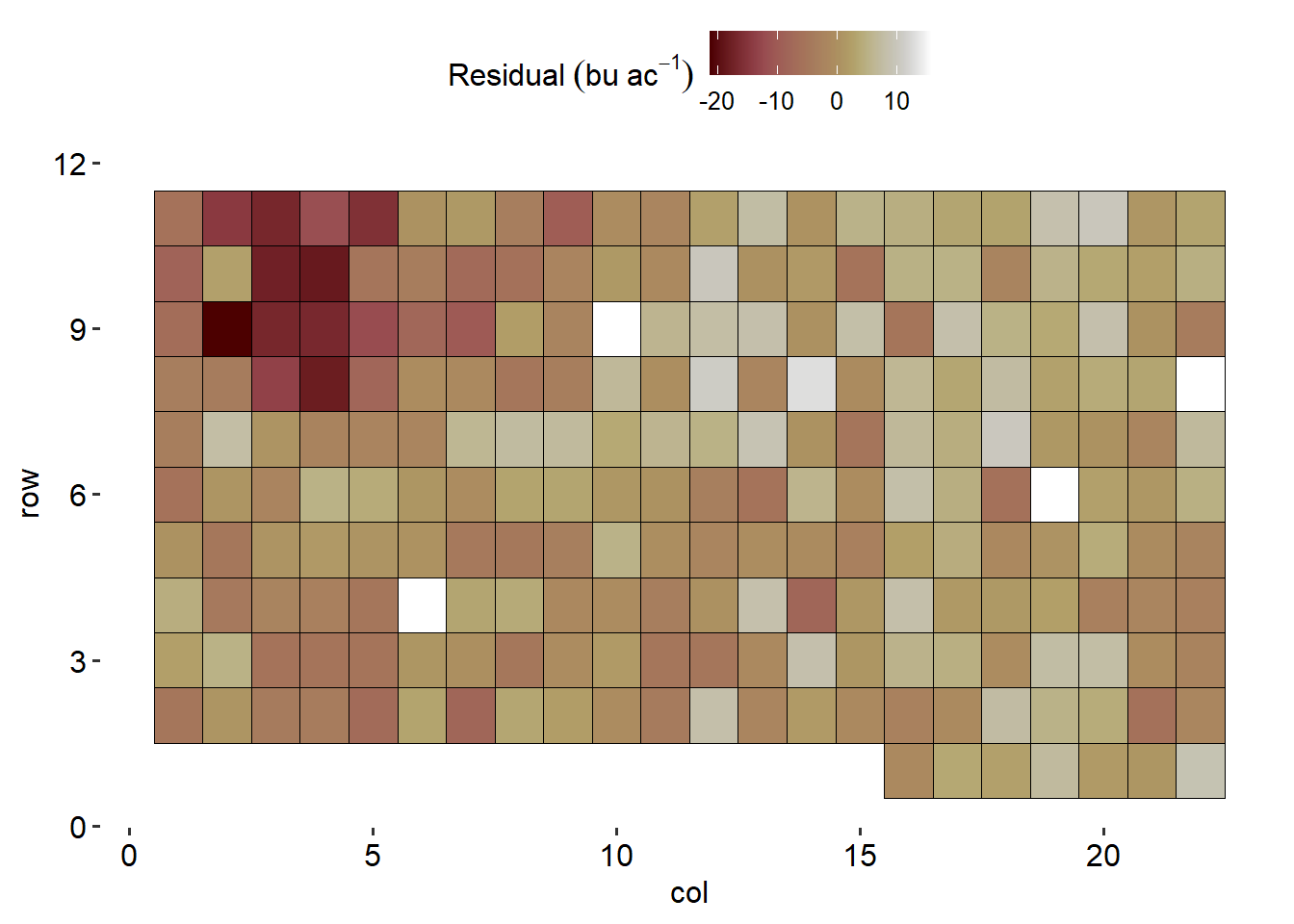
## gen emmean SE df lower.CL upper.CL
## NE86503 32.6500 3.855689 66.69 24.95335 40.34665
## NE87619 31.2625 3.855689 66.69 23.56585 38.95915
## NE86501 30.9375 3.855689 66.69 23.24085 38.63415
## Redland 30.5000 3.855689 66.69 22.80335 38.19665
## Centurk78 30.3000 3.855689 66.69 22.60335 37.99665
## NE83498 30.1250 3.855689 66.69 22.42835 37.82165
##
## Degrees-of-freedom method: kenward-roger
## Confidence level used: 0.95library(glmmTMB)
data$coordinate <- numFactor(data$col, data$row)
data$group <- factor(1)
m_spatial <- glmmTMB(yield ~ gen + (1|rep) + mat(coordinate + 0 | group), data = data)data$resid_spatial <- resid(m_spatial)
data %>%
ggplot(aes(col, row))+
geom_tile(aes(fill = resid_spatial), color = "black")+
theme_pubr()+
scico::scale_fill_scico()+
theme(axis.line = element_blank())+
labs(fill = expression(Residual~(bu~ac^{-1})))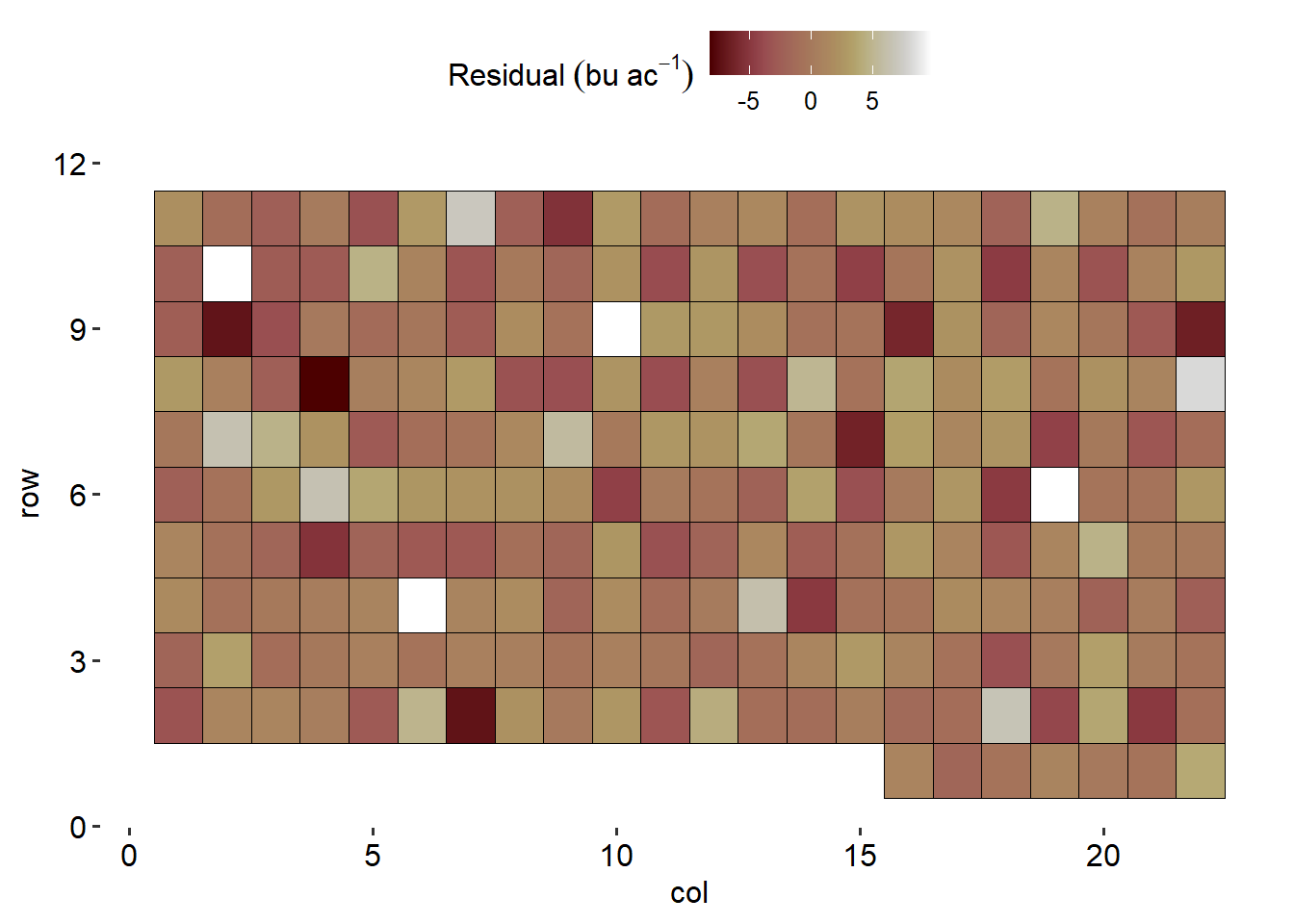
## gen emmean SE df asymp.LCL asymp.UCL
## Buckskin 33.37166 3.399255 Inf 26.70925 40.03408
## NE85556 29.35020 3.395673 Inf 22.69480 36.00560
## NE87619 29.09390 3.397032 Inf 22.43584 35.75196
## NE83498 28.90073 3.405042 Inf 22.22697 35.57449
## Redland 28.63517 3.393941 Inf 21.98317 35.28717
## NE86503 27.51328 3.451773 Inf 20.74792 34.27863
##
## Confidence level used: 0.95